|
Xpl<-xp_hat + qnorm(0.025)*sqrt(v_xp_hat) #lower bound Xp_hat<-fd$estimate+qprobs*fd$estimate #estimated perc. To add the confidence bounds as in Minitab, you can do the following fd<-fitdistr(x, "normal") #Maximum-likelihood Fitting of Univariate Dist from MASS Ggplot(data = df, aes(x = x, y = y)) + geom_point() + geom_abline(intercept = int,slope = slope)+scale_y_continuous(limits=range(qprobs), breaks=qprobs, labels = 100*probs)+labs(y ="Percent", x="Data") The following shows the line in the right spot: dfI've learned that the last two rows of the legend refer to the Anderson-Darling test for normality and can be reproduced with the nortest package. I get similar results using R's base graphics plot(df$x, df$y) Versus: ggplot(data = df, aes(x = x, y = y)) + geom_point() + geom_abline(intercept = int, slope = slope) However, incorporating this information into ggplot2 or base graphics does not yield the same results. Indeed, comparing these results to what you get out of the probplot object seem to compare very well: > check str(check) Sample data to recreate the plot above: x xl yl slope int slope  It seems that geom_smooth() would be the likely candidate to add the bands, but I haven't figure that out.įinally, the Getting Genetics Done guys describe something similar here. Similarly, ggplot's stat_qq() seems to present similar information with a transformed x axis. 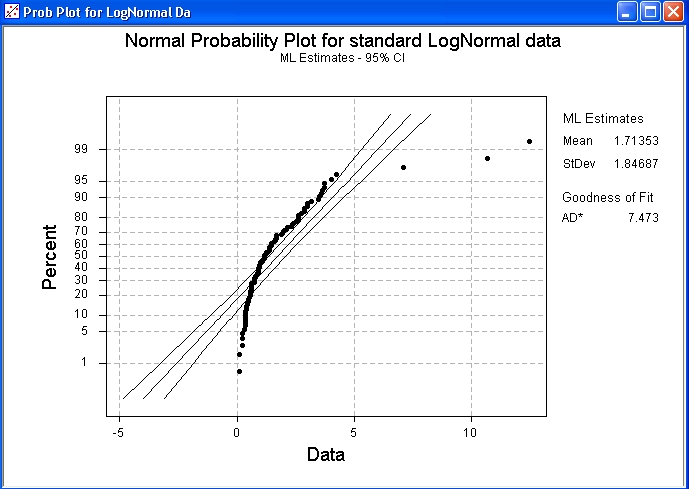 Unfortunately, I cannot figure out how to add the confidence interval bands around this plot. The probplot gets you most of the way there. Minitab describes this as a normal probability plot. I am trying to recreate the following plot with R.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
